
Features
Agronomy
Diseases
Into the deep
Science is about asking questions, and trying to answer them. Of course, some are easier to answer than others. With regards to potato diseases of the soil, questions have been hard to answer indeed. “The soil is an extremely complex ecological environment,” observes molecular biologist Dr. Claudia Goyer of Agriculture and Agri-Food Canada’s Potato Research Centre in Fredericton, N.B. “In a tiny amount of soil, there are millions of organisms of many types, and on the whole, our knowledge of them is very incomplete or completely nonexistent.”
The limited knowledge that we currently have about soil organisms and interactions between them is what makes the creation and availability of new technologies so important. New devices and strategies – in this case, a “deep sequencer” and its associated powerful DNA analysis programs – can help scientists get answers to questions that were difficult or impossible to answer before. “In simple terms, this will enable us to find out what organisms and how many of them are present in the soil through looking at their DNA, which will in turn enable us to better help the soil itself battle the diseases that affect our crops,” says Goyer.
Goyer explains that while we already know that manipulating soil conditions through changes in agricultural practices can reduce or prevent disease, up until recently, we haven’t had an effective way of gauging how well we are doing. “We know that by doing things like adding compost or manure, we can create conditions that reduce or increase disease,” Goyer notes, “but we haven’t had a way to get a good handle on how soil microbial communities are changing and how these changes influence plant pathogens. It is also not clear, for example, how much compost we need to add to reduce a soil-based plant pathogen, or how other factors such as soil type, moisture or pH might also affect the desired outcome.”
While the global scientific community has already had the capacity to isolate and sequence DNA from organisms for a long time, soil has proven a greater challenge. “The amount of various organisms in the soil is so large, that a much more powerful sequencing and analysis tool was required,” Goyer says. “Now we have it. This is a huge step forward technologically. Now we can sequence so much more and we have a lot more computing power at our fingertips to sort it all out in terms of which sequences belong to which organisms and how many of them are present.” The deep sequencer identifies millions of pieces of genetic data. The software then assembles the genetic pieces, makes comparisons to organisms already known, and points out unknowns.
“Once you know what is there and how much of certain organisms are there, you can also answer other questions about the functioning of the soil,” says Goyer, “such as why certain organisms are there in the first place, and why others aren’t.”
Deep sequencing has been around about nine years, but about five years ago, the cost became more reasonable. “This year, we are going to be using large deep sequencers in Quebec or Ontario, but there are smaller deep sequencers now available and used in many labs around the world,” Goyer notes. “The smaller versions have a smaller capacity, but are suitable for some tasks. You can use smaller or larger tech, depending on what you want to do.” Goyer explains that it would only cost about $1,000 to sequence something the size of the human genome at this point in time, whereas years ago, it was many orders of magnitude more expensive.
Control of common scab
Goyer recently discovered a Bacillus bacterium in the soil that inhibits the growth of another bacterium called Streptomyces scabies, the cause of common scab. Once common scab is in the soil, it’s difficult to reduce or eliminate the pathogen population. “In terms of controlling or getting rid of Streptomyces scabies, we may be able to add more of a certain organism like this Bacillus bacterium,” Goyer notes. “It’s found naturally in soil along with lots of other ‘good’ bacteria. We also want to discover if the presence of other microbes is conducive to having common scab, to the growth of Streptomyces scabies.”
She says now that deep sequencing is available, it can also be used to see how changing agricultural practices such as rotation or application of compost will affect how many Streptomyces are present and how the microbial communities are changing. Goyer adds, “It’s a much more precise look at what is happening with soil organisms. It’s very exciting because it allows us to do experiments we couldn’t do before, because there was no effective technology to get these results.”
In 2012, the Potato Research Centre marked its 100th birthday. Back then, soil analysis involved dissolving a sample in liquid, shaking it and seeing what they could observe with the eye and under the microscopes available at the time. Deep sequencing, in comparison, is leagues ahead. “It’s the latest in a number of technologies that have come along over the years and it provides us with more genetic information than ever before,” Goyer says. “With it, scientists like us will continue to find all kinds of new life and be able to answer questions that we could only dream of answering decades ago. It’s a very great time to be a soil microbiologist.”
November 19, 2013 By Treena Hein
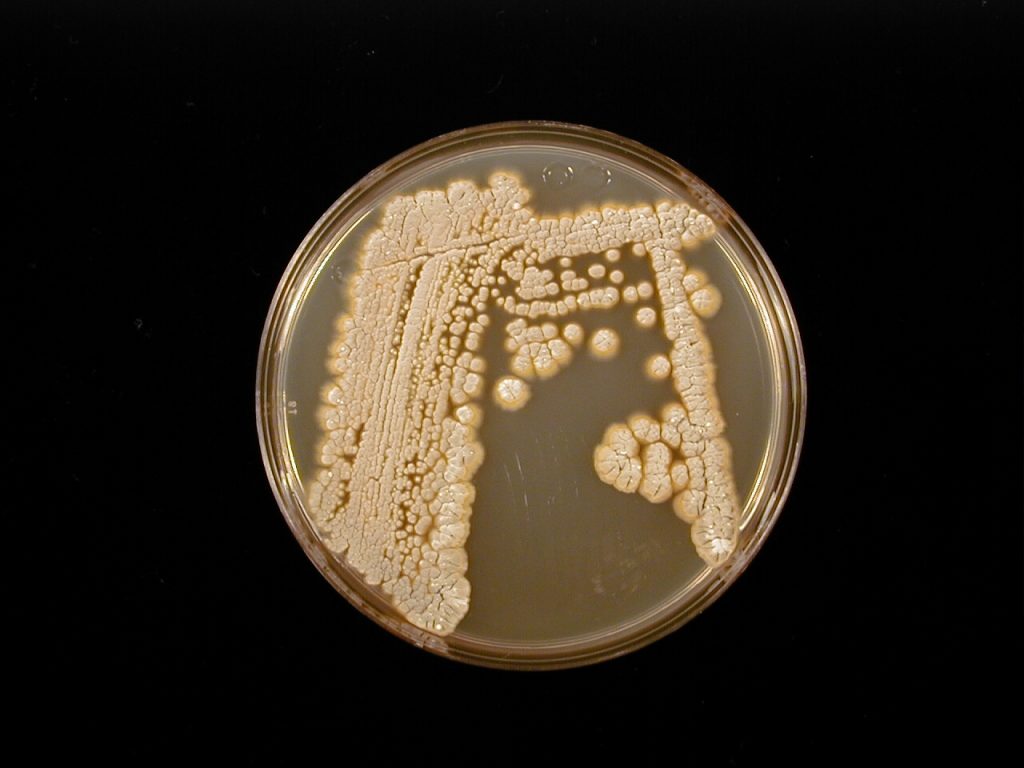 The common scab pathogen named Streptomyces scabies With regards to potato diseases of the soil
The common scab pathogen named Streptomyces scabies With regards to potato diseases of the soilPrint this page